An Elliptic Kernel Unsupervised Autoencoder-Graph Convolutional Network Ensemble Model for Hyperspectral Unmixing
Authors
Estefania Alfaro-Mejia, Carlos J Delgado, Vidya Manian
Abstract
Spectral Unmixing is an important technique in remote sensing used to analyze hyperspectral images to identify endmembers and estimate abundance maps. Over the past few decades, performance of techniques for endmember extraction and fractional abundance map estimation have significantly improved. This article presents an ensemble model workflow called Autoencoder Graph Ensemble Model (AEGEM) designed to extract endmembers and fractional abundance maps. An elliptical kernel is applied to measure spectral distances, generating the adjacency matrix within the elliptical neighborhood. This information is used to construct an elliptical graph, with centroids as senders and remaining pixels within the geometry as receivers. The next step involves stacking abundance maps, senders, and receivers as inputs to a Graph Convolutional Network, which processes this input to refine abundance maps. Finally, an ensemble decision-making process determines the best abundance maps based on root mean square error metric. The proposed AEGEM is assessed with benchmark datasets such as Samson, Jasper, and Urban, outperforming results obtained by baseline algorithms. For the Samson dataset, AEGEM excels in three abundance maps: water, tree and soil yielding values of 0.081, 0.158, and 0.182, respectively. For the Jasper dataset, results are improved for the tree and water endmembers with values of 0.035 and 0.060 in that order, as well as for the mean average of the spectral angle distance metric 0.109. For the Urban dataset, AEGEM outperforms previous results for the abundance maps of roof and asphalt, achieving values of 0.135 and 0.240, respectively. Additionally, for the endmembers of grass and roof, AEGEM achieves values of 0.063 and 0.094.
Concepts
The Big Picture
Imagine looking down at a forest from a satellite. What you see isn’t a clean mosaic of trees, water, and soil. It’s a blurry soup of mixed signals. Each pixel captured by the sensor might contain a little bit of everything: some tree canopy, a patch of wet ground, a glint of rooftop. Figuring out exactly what percentage of each pixel is “tree” versus “water” versus “asphalt” is one of remote sensing’s oldest and most stubborn problems.
This is the challenge of hyperspectral unmixing: taking a rich, multi-wavelength image and decomposing it into its pure material constituents (called endmembers) along with maps showing how much of each material lives in every pixel (abundance maps). Traditional approaches either oversimplify by assuming each pixel contains only one material, or they struggle with the spatial complexity of real terrain. Deep learning has improved things, but large, sprawling features like rooftops, water bodies, and tree stands remain hard to reconstruct accurately.
A team from the University of Puerto Rico’s AI Imaging Group introduced AEGEM, the Autoencoder Graph Ensemble Model, which stitches together three ideas into a single pipeline that outperforms existing methods on multiple benchmark datasets.
Key Insight: By combining an unsupervised autoencoder with a graph convolutional network guided by an elliptical neighborhood geometry, AEGEM captures both the spectral fingerprints and the spatial relationships of materials simultaneously, then uses ensemble decision-making to pick the best result.
How It Works
The AEGEM pipeline runs in eight stages, but the core logic unfolds in three acts.
Act 1: The autoencoder extracts initial endmembers and abundance maps. A convolutional autoencoder (a neural network that compresses data to a bottleneck and reconstructs it) learns to separate the hyperspectral image into its constituent materials without any labeled training data. This is purely unsupervised learning: no one tells the network what “water” or “tree” looks like. The encoder estimates abundance maps; the decoder handles endmember extraction and image reconstruction.

Act 2: An elliptical kernel builds a spatial graph. Once the autoencoder produces initial abundance estimates, the model applies an elliptical kernel to compute spectral distances between neighboring pixels. Why elliptical? Real features like fields and roads tend to stretch in one direction rather than spreading out as perfect circles. An ellipse captures that directionality better than a simple circular or square grid.
The key steps in building this graph:
- Select centroid pixels as graph nodes
- Apply the elliptical kernel to compute spectral distances within each neighborhood
- Assign directional edges from centroids (senders) to surrounding pixels (receivers)
- Stack the abundance maps, sender features, and receiver features as input to the next stage
Act 3: A graph convolutional network refines the maps, and an ensemble picks the winner. The stacked inputs feed into a Graph Convolutional Network (GCN), which propagates information along the edges of the spatial graph to refine the abundance estimates. GCNs are built for this kind of message-passing: each node updates its representation by aggregating information from its neighbors. AEGEM runs this whole process across multiple configurations and applies a root mean square error (RMSE) ensemble criterion, selecting the result with the smallest average prediction error.
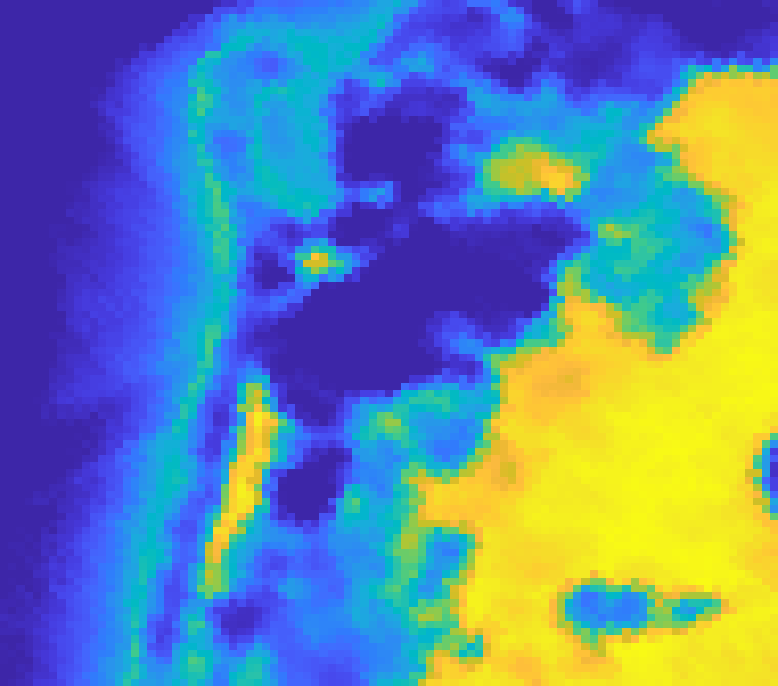
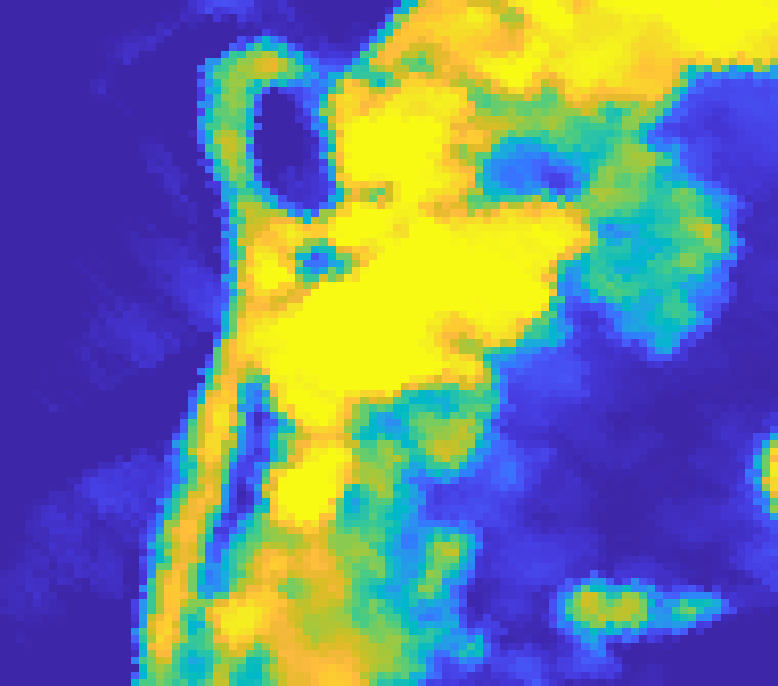
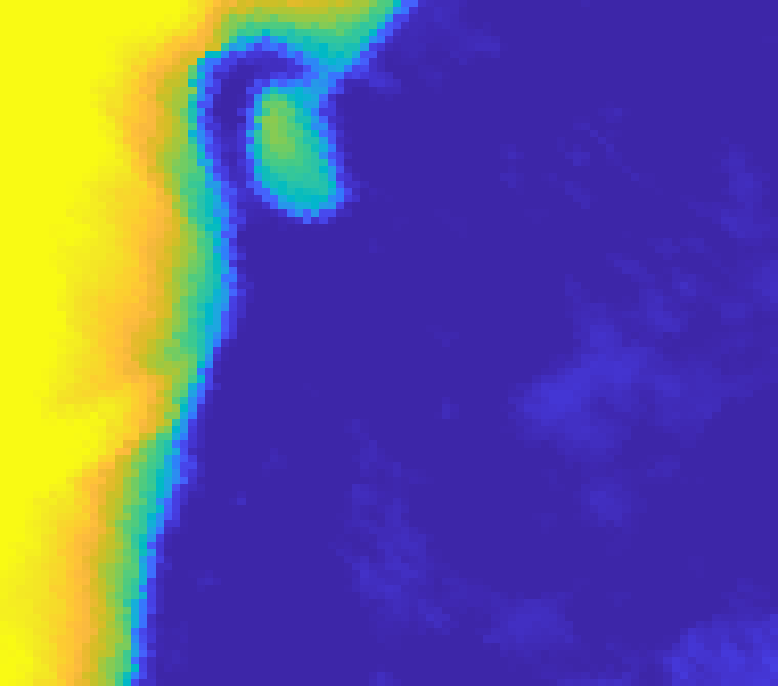
On the Samson dataset (a classic benchmark with water, tree, and soil classes) AEGEM achieves RMSE values of 0.081 (water), 0.158 (tree), and 0.182 (soil), beating all baselines across all three classes. On the Jasper dataset, tree and water endmembers score 0.035 and 0.060, with a mean spectral angle distance of 0.109. The challenging Urban dataset is where things get most competitive: AEGEM outperforms on roof (0.135) and asphalt (0.240) abundance maps, while also achieving top scores for grass endmembers (0.063) and roof endmembers (0.094).

Why It Matters
Hyperspectral unmixing feeds directly into precision agriculture (identifying crop stress before it’s visible to the naked eye), environmental monitoring (tracking land cover change over time), urban planning, and medical imaging. Accurately decomposing mixed signals into pure components is a fundamental data problem, and the techniques that solve it in remote sensing often transfer well to other domains.
Beyond the benchmark numbers, AEGEM’s architectural philosophy is what stands out. Treating pixels not as isolated points but as nodes in a spatially-aware graph means the model explicitly encodes the structure of the scene. The elliptical kernel is a smart design choice: spectral neighborhoods don’t spread out uniformly in every direction. Materials don’t distribute themselves in perfect circles. The ellipse is a small but meaningful built-in assumption that better matches the geometry of real terrain, the kind of domain-informed design that tends to generalize well.
Where could this go next? Adaptive kernels that learn their own shape. Multi-scale graph hierarchies for larger spatial extents. Change-detection applications where unmixing accuracy determines whether subtle land-cover shifts get noticed at all.
Bottom Line: AEGEM shows that combining autoencoders, graph networks, and geometric kernels into a principled ensemble pipeline can squeeze significantly better abundance maps out of hyperspectral data, a win for anyone who needs to know what the Earth’s surface is actually made of, pixel by pixel.
IAIFI Research Highlights

This work connects machine learning architecture design with the physical realities of spectral remote sensing, using elliptical geometry to encode how materials actually distribute themselves across terrain rather than imposing arbitrary spatial assumptions.

AEGEM introduces an ensemble strategy for fusing autoencoder and graph convolutional outputs, pushing forward unsupervised representation learning for structured spatial data.

More accurate decomposition of mixed spectral signals into pure material components improves monitoring of Earth systems, from vegetation health to urban heat islands, where precise material-level analysis matters.
Future directions include adaptive kernel learning and multi-scale graph hierarchies for larger regions of interest; the work comes from the AIIG Laboratory at the University of Puerto Rico, Mayaguez, with NASA funding (grant 80NSSC21M0155). [arXiv:2406.06742](https://arxiv.org/abs/2406.06742)
Original Paper Details
An Elliptic Kernel Unsupervised Autoencoder-Graph Convolutional Network Ensemble Model for Hyperspectral Unmixing
[arXiv:2406.06742](https://arxiv.org/abs/2406.06742)
Estefania Alfaro-Mejia, Carlos J Delgado, Vidya Manian

