Reinforcement Learning for Control of Non-Markovian Cellular Population Dynamics
Authors
Josiah C. Kratz, Jacob Adamczyk
Abstract
Many organisms and cell types, from bacteria to cancer cells, exhibit a remarkable ability to adapt to fluctuating environments. Additionally, cells can leverage a memory of past environments to better survive previously-encountered stressors. From a control perspective, this adaptability poses significant challenges in driving cell populations toward extinction, and thus poses an open question with great clinical significance. In this work, we focus on drug dosing in cell populations exhibiting phenotypic plasticity. For specific dynamical models switching between resistant and susceptible states, exact solutions are known. However, when the underlying system parameters are unknown, and for complex memory-based systems, obtaining the optimal solution is currently intractable. To address this challenge, we apply reinforcement learning (RL) to identify informed dosing strategies to control cell populations evolving under novel non-Markovian dynamics. We find that model-free deep RL is able to recover exact solutions and control cell populations even in the presence of long-range temporal dynamics. To further test our approach in more realistic settings, we demonstrate robust RL-based control strategies in environments with measurement noise and dynamic memory strength.
Concepts
The Big Picture
Imagine trying to exterminate a colony of ants. You spray the nest, most die, but a few survivors seem to “remember” the attack. The next time you spray, they’re ready. Their offspring inherit that readiness.
Now scale this to billions of cancer cells inside a human body, and the stakes become obvious.
This is the central challenge of cancer chemotherapy and antibiotic treatment. Cell populations don’t just evolve resistance passively over generations. They switch between vulnerable and resistant states in real time, driven by the very drugs meant to kill them. Worse, cells can encode molecular memory of past drug exposure, staying primed to resist the next round even after treatment ends. Traditional strategies (dose hard, dose constantly) often backfire, selecting for the most resistant survivors.
Josiah Kratz and Jacob Adamczyk tackled this directly. They built a mathematical model for memory-driven cell behavior, then trained an AI through trial and error to discover dosing strategies that can drive even these adaptive, history-aware populations toward extinction.
Key Insight: When cancer cells and bacteria “remember” past drug exposure, optimal treatment can’t be designed from the cell’s current state alone, but an AI trained on short snapshots of recent history can find effective dosing strategies that work even when the underlying biology is unknown.
How It Works
The foundation is a phenotypic switching model, a mathematical description of how cells toggle between a drug-vulnerable state and a drug-resistant state. When drug is applied, vulnerable cells die but also switch to a resistant form; when drug is removed, resistant cells gradually revert to vulnerability. Some resistant cells actually grow fastest in the presence of the drug, a phenomenon called drug addiction that has been observed in real cancer cell lines.
The distinguishing feature here is non-Markovian dynamics: the cell population’s future behavior doesn’t depend only on its current state, but on its entire history of drug exposure. The authors model this with a memory kernel, a mathematical function that weights how strongly past drug concentrations influence current switching rates. A population with long-range memory responds to today’s dose differently depending on whether it was treated heavily last week or not at all. This is far more realistic than standard models, but it makes the control problem exponentially harder.
For systems with known parameters and simple (Markovian) dynamics, the exact optimal strategy can be derived analytically. It turns out to be bang-bang control: give the maximum dose or zero dose, never anything in between. The researchers prove this theoretically and use it as a hard constraint for their AI agent. When parameters are unknown or memory dynamics are at play, that exact solution is out of reach.
This is where deep reinforcement learning (RL) comes in. The agent learns by trial and error, discovering which actions lead to good outcomes through millions of simulated attempts. The pipeline works in three steps:
- Observe: The agent sees current counts of vulnerable and resistant cells.
- Stack: Rather than the current state alone, the agent receives a short window of recent observations, a technique called framestacking borrowed from video game AI.
- Act: The agent selects a dose, is rewarded for reducing total cell count, and penalized for population growth.
Framestacking is what makes memory tractable. The agent infers the system’s history from recent observations without needing to know the underlying model parameters. That’s exactly the situation a clinician faces when treating an unknown tumor.
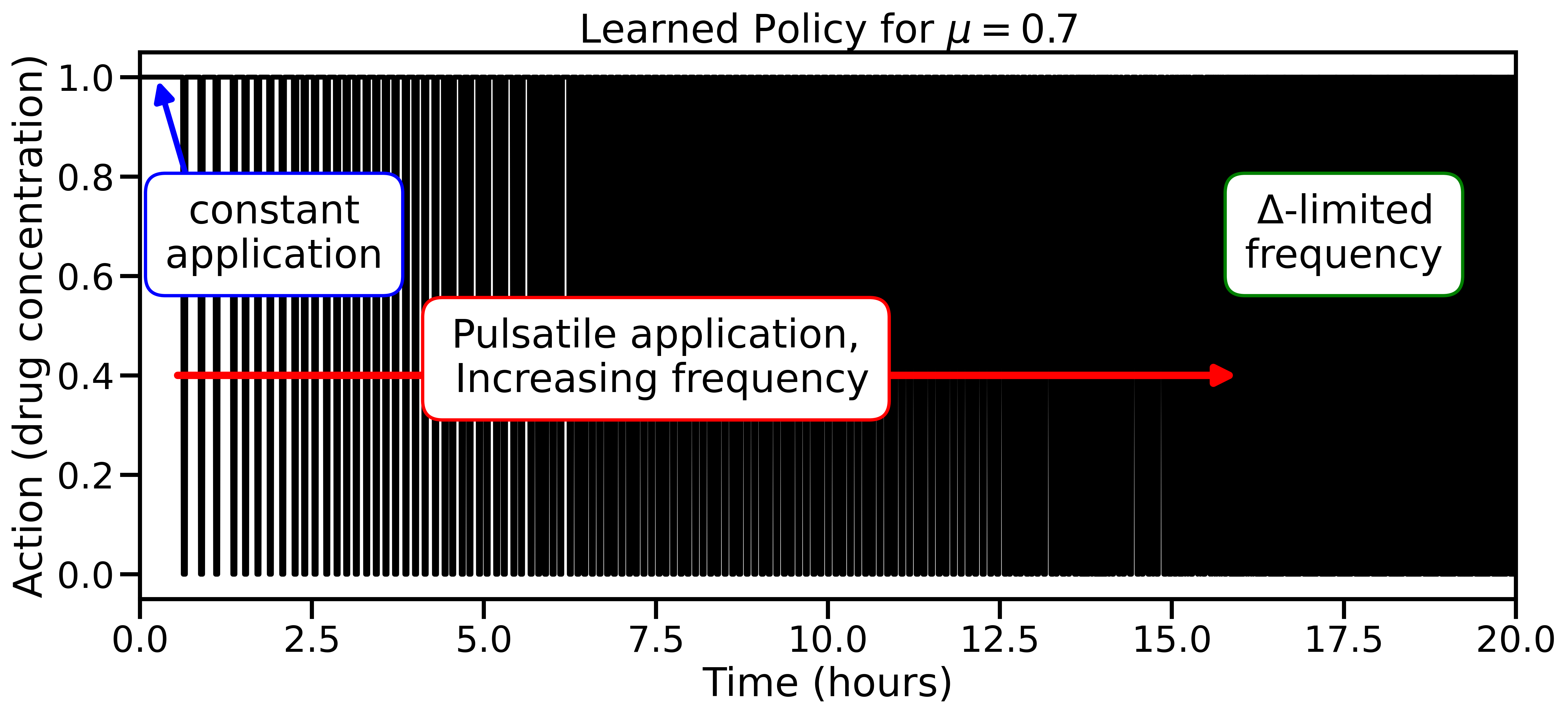
Why It Matters
Across Markovian models where exact solutions are known, the RL agent recovers them independently. That’s a clean validation that the approach is sound.
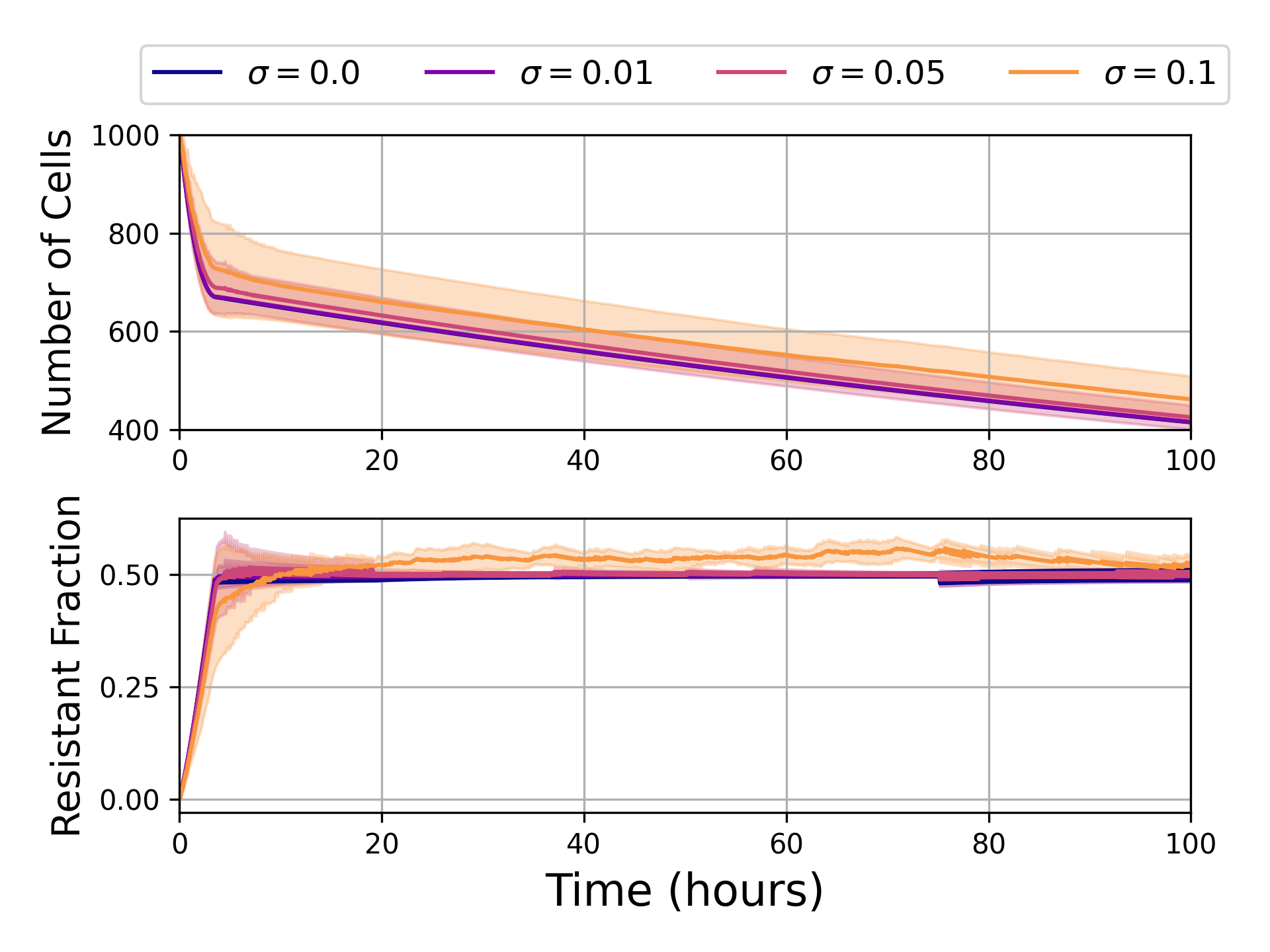
The real test is the non-Markovian regime, where no exact solution exists. Here, framestacked RL still finds strong control policies as memory strength varies and even when observations are corrupted by realistic measurement noise. The agent learns a single policy that doesn’t need retraining for different memory parameters, which is clinically essential since you rarely know a tumor’s exact molecular memory profile.
This work sits at a productive intersection of physics and machine learning. The authors bring tools from optimal control theory, including bang-bang constraints and Pontryagin’s maximum principle, to guide what would otherwise be a pure black-box learning problem. The result is an agent that doesn’t just stumble onto a good policy; it finds policies that respect the underlying physics.
That combination of principled modeling and data-driven optimization is increasingly how the hardest problems in biophysics and medicine get solved. The framework extends beyond cancer: bacterial biofilms, immunotherapy resistance, even ecosystems responding to environmental change are all adaptive systems with memory that could benefit from this approach.
Next steps are clear. The authors plan to move from deterministic population models to stochastic ones, incorporate pharmacokinetic effects (how drugs actually distribute and decay in the body), and eventually connect to real experimental data from cell culture or clinical trials.
Bottom Line: By combining a memory-based cell-switching model with deep RL and framestacking, Kratz and Adamczyk show that AI can find near-optimal drug dosing strategies even when cells “remember” past treatment, a concrete step toward genuinely adaptive, personalized cancer therapy.
IAIFI Research Highlights

This work fuses optimal control theory from physics with model-free deep reinforcement learning from AI to solve a biologically realistic dosing problem that neither field could crack alone.

Incorporating physics-derived constraints (bang-bang control) into RL training sharply improves sample efficiency and policy quality, offering a blueprint for physics-informed RL in complex dynamical systems.

The paper introduces a non-Markovian phenotypic switching model that captures long-range temporal memory in biological populations, advancing the mathematical physics of living adaptive systems.
Future work will extend the framework to stochastic dynamics and experimental validation; the paper was presented at ICLR 2025 and is available from IAIFI-affiliated author Jacob Adamczyk at UMass Boston.
Original Paper Details
Reinforcement Learning for Control of Non-Markovian Cellular Population Dynamics
2410.08439
["Josiah C. Kratz", "Jacob Adamczyk"]
Many organisms and cell types, from bacteria to cancer cells, exhibit a remarkable ability to adapt to fluctuating environments. Additionally, cells can leverage a memory of past environments to better survive previously-encountered stressors. From a control perspective, this adaptability poses significant challenges in driving cell populations toward extinction, and thus poses an open question with great clinical significance. In this work, we focus on drug dosing in cell populations exhibiting phenotypic plasticity. For specific dynamical models switching between resistant and susceptible states, exact solutions are known. However, when the underlying system parameters are unknown, and for complex memory-based systems, obtaining the optimal solution is currently intractable. To address this challenge, we apply reinforcement learning (RL) to identify informed dosing strategies to control cell populations evolving under novel non-Markovian dynamics. We find that model-free deep RL is able to recover exact solutions and control cell populations even in the presence of long-range temporal dynamics. To further test our approach in more realistic settings, we demonstrate robust RL-based control strategies in environments with measurement noise and dynamic memory strength.

